By Stephanie Norman, DVM, MS, PhD | Marine Mammal Vet for Central Puget Sound MMSN
Do Pacific Northwest marine mammals carry antibiotic-resistant bacteria from land animals?
Antibiotic resistance, a global concern, is a significant health issue of animals and humans. Resistant bacteria, a growing presence in marine life, are derived from land via humans, animals, and agriculture. Antibiotic resistance is documented in multiple marine species. Data on resistance in marine species in Washington State's inland waters, known as the Salish Sea, is relatively limited.
Preliminary work reported resistance in young harbor seals (Phoca vitulina) in rehabilitation and in breath from local southern resident killer whales (Orcinus orca), as well anecdotal reports from stranding necropsies. A larger dataset with multiple species, age classes, and locales is warranted to determine if resistant bacteria exhibit patterns within this urban marine ecosystem, and if they pose a threat to other marine mammals and human health. So we set out to answer the question: What is the prevalence of resistant bacteria present in marine mammals of an urban ecosystem, the Salish Sea, in Washington State?
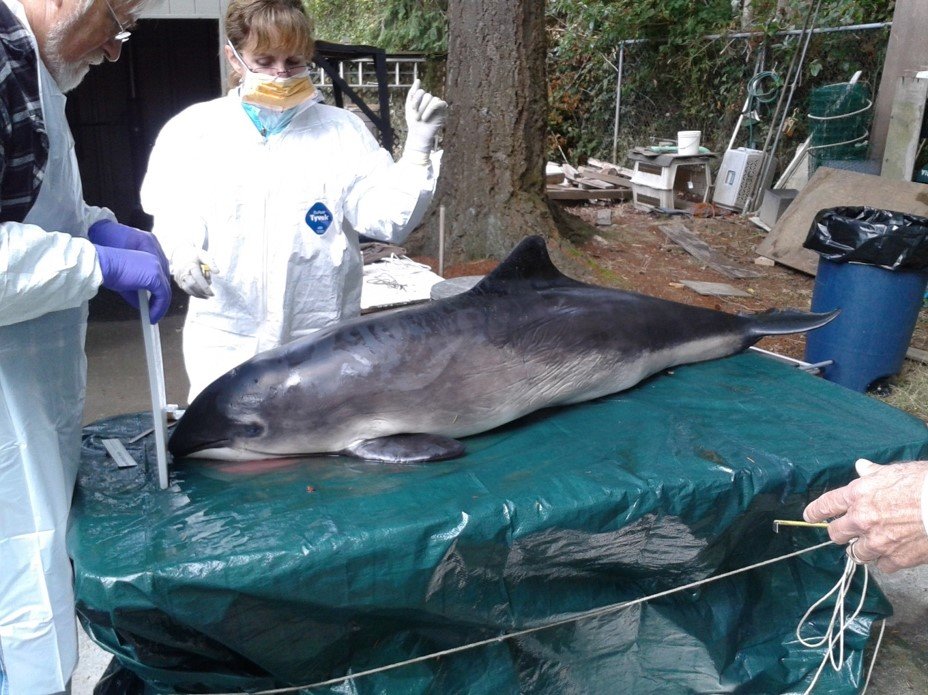
This crowdfunded and Wildlife Disease Association-supported project seeks to: detect and describe the presence and distribution of antibiotic resistance in two key Salish Sea marine mammal species; determine differences in resistance between harbor seals and harbor porpoises (Phocoena phocoena); and document geographic patterns. These species were selected because they are abundant and relatively site-specific within the Salish Sea. A cross-sectional sampling study is being conducted to calculate prevalence (%) of resistant bacteria from 69 dead-stranded harbor seals and 69 porpoise, of all age classes and sexes. The study area is divided into northern and southern sampling regions. We hypothesize that resistance prevalence will: be < 5% overall, differ between harbor seals and porpoises and between the two sampling regions. Sterile swabs of the intestines and lesions noted on necropsy are submitted for aerobic culture and sensitivity to a suite of 16 commonly used antibiotics using standard laboratory techniques.
Samples have been collected from ten harbor seals and 5 harbor (Table 1). Preliminary results (% of each species sampled to-date) indicate multi-drug resistant bacteria were detected in four seals (40%) and two porpoises (40%), respectively, with all age classes and sexes involved (see table attached at the end of this report). Swabs for culture and sensitivity were submitted by Central Puget Sound Marine Mammal Stranding Network (n=1), Washington Department of Fish and Wildlife (n=1), Cascadia Research Collective (n=4), The Whale Museum/San Juan County Marine Mammal Stranding Network (n=3), Sno-King Marine Mammal Response (n=2), Seal Sitters (n=1) and Port Townsend Marine Science Center/East Jefferson County Marine Mammal Stranding Network (n=1).
Genotyping and sequencing of E. coli cultures are ongoing. Study results will better define antibiotic resistance prevalence patterns and risk factors in Salish Sea marine mammals. In addition to Orca Network and Central Puget Sound Marine Mammal Stranding Network, project partners include Marine-Med , Washington State Department of Wildlife , Cascadia Research Collective, The Whale Museum/San Juan County Marine Mammal Stranding Network, Whatcom County Marine Mammal Stranding Network, World Vets, Port Townsend Marine Science Center, and Sno-King Marine Mammal Response. Initial support for this project was through Experiment.com , with additional support by the Wildlife Disease Association.
YOUR SUPPORT through GlobalGiving provides additional funding for Dr. Norman's work to carry out this project, as well as her work with the Central Puget Sound Marine Mammal Stranding Network to respond to and investigate marine mammal strandings, perform necropsies and conduct research on the health of marine mammals of the Salish Sea.
Links:
Project reports on GlobalGiving are posted directly to globalgiving.org by Project Leaders as they are completed, generally every 3-4 months. To protect the integrity of these documents, GlobalGiving does not alter them; therefore you may find some language or formatting issues.
If you donate to this project or have donated to this project, you can receive an email when this project posts a report. You can also subscribe for reports without donating.